Raster3D import filters
grasp/ungrasp
Grasp (home page)
is a molecular visualization tool that to the best of my knowledge is available
only for SGI workstations and some older versions of Windows. Grasp itself does
not produce Raster3D output, but the molecular surfaces calculated
by grasp can be saved to a file and then converted to a Raster3D input
description. The filter program that does this conversion is called
"ungrasp", and is part of the Raster3D source distribution. The
ungrasp source code and notes on using it are in the ./grasp/ subdirectory
of the Raster3D source tree.
MSMS
MSMS
(original home page)
is another program that calculates molecular surfaces, and has a number of advantages over
grasp. Although you may still be able to download MSMS itself, it has been replaced by
a newer python-based tool PMV
(home page).
Two filter programs to convert MSMS output files into Raster3D input files are
provided in the Raster3D source distribution. One is a perl script. The other is C source
code. Both are in the ./examples/msms subdirectory of the source tree, and by default
are installed permanently in /usr/local/share/Raster3D/examples/msms/.
| A figure composed with MSMS and Raster3D |
The script that generated this figure |
|---|
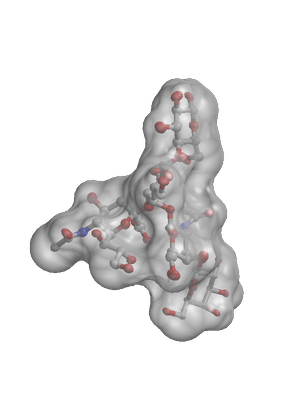 |
#!/bin/csh
#
# Calculate molecular surface of a pentasaccharide
#
ln -s ../sugarG.pdb protein.pdb
cat protein.pdb | pdb_to_xyzr > surf.dat
msms -if surf.dat -of surf -probe_radius 1.4 -density 5.0
#
# Convert msms output to Raster3D description of the surface
#
./msms2r3d.pl surf.face surf.vert > surf.r3d
#
# Make an auto-centered ball+stick view of the molecule
#
rastep -auto -rad 0.20 -prob 0.10 < protein.pdb > protein.r3d
#
# Combine the ball+stick figure with a transparent material
# description from the Raster3D library
# applied to the molecular surface generated in MSMS
# Save the combined picture as a PNG file
#
render -size 300x400 -zoom 85% -png picture.png << EOF
@protein.r3d
@translucent_60
@surf.r3d
EOF
#
rm -f protein.pdb msms_surf.dat.log
rm -f surf.dat surf.face surf.vert
rm -f surf.r3d protein.r3d
|
 Back to top
Back to top
 Raster3D homepage
Raster3D homepage

 Back to top
Back to top